The right answers to frequently asked questions
Find the answers to all our products and services by clicking the links below.
What Species can be detected using Microbiome Profiling by NGS?
Depending on your choice of target regions, all bacterial, archeal and fungal species of which a sequence is published in the NCBI nucleotide database can be identified. There are currently for example thousands of bacterial species with sequence entries in the database. Closely related species with almost identical DNA sequences will not be discriminated and only the genus (or the family) will be reported.
What are the limitations of using Microbiome Profiling by NGS?
• Closely related species are not distinguishable because of sequence identity.
• Degradation of bacterial DNA may occur in highly processed samples or samples with low pH - a species identification is therefore difficult and sometimes not possible.
• The more starting material we receive, the higher the probability to get excellent sequencing results.
What is the level of false positives / negatives?
• Samples with strongly degraded DNA will give the result “no species identification possible”. To avoid false-positive results, a negative control (water) is always co-analysed with the samples.
• With NGS all DNA amplicons that were successfully amplified from the sample DNA will be detected. Due to the high sensitivity of the method, sequences with low presence in the sequence pool or sequences with unexpected sequence lengths have to be removed from the data analysis during the bioinformatical workflow in order to exclude false-positives due to PCR and/or sequencing errors. Species with a fraction below 0.5 % of all assigned reads are not reported.
How should I choose the target regions and the amount of targets to be analysed?
For bacterial profiling please choose at least on 16S target, for fungal profiling at least one ITS target. For taxonomic profiling or archea please selcet our archeal 16S target.
As each primer set fails to amplify certain species we recommend that you carefully check the literature available about the limitations of each primer set. It is recommended to use multiple primer combinations to capture the entire community (e.g. Ihrmark et al., 2012; Toju et al., 2012).
What quality control do you apply on my samples?
We use the amplicon generation process as the entry quality control. Even though the PCR conditions are optimized the success of the amplicon generation depends on the amount of bacterial, archaeal or fungal DNA in the sample. In case we can not generate an amplicon or only a very week amplicon this is a strong indication that the content was too low. By default we will repeat the amplicon generation once with modified conditions.
What happens if no or only a weak amplicon can be generated from some of my samples:
In case no amplicon can be generated, we will stop processing of those samples and reimburse you for the sequencing part that was not performed. In the event that only a weak amplicon can be generated, Eurofins Genomics will include those amplicons in the amplicon pool for sequencing, too. Experience shows that those amplicons will result in very few reads only and interpretation of those reads should be done with special care.
When choosing your service on MiSeq, can we send samples that consist of several amplicons generated with different primer pairs?
Usually you would send per sample 1 amplicon generated with 1 primer pair. In some cases you might require however less reads per amplicon as given in the Project Specification. In such cases the individual amplicon samples you send may also consist of several amplicons. Please note, that in this case however we will not perform the clipping of primer sequences and deliver raw data only.
How should I send my samples?
- Please use 1.5 ml safe-lock tubes for your templates and primers
- Do not tape or wrap tubes with parafilm. Safe-lock tubes offer perfect sealing and evaporation protection
- Label your template with our free NGS barcode labels.
If you send more than 92 samples then you can also sent the samples in plates. Please use full skirted PCR plates (e.g. Eppendorf or Twin).
For more information and for sample submission for extraction please read our Sample Submission Guide.
(Picture shows the procedure with Sanger Prepaid Barcodes but work the same with free NGS Barcodes)
|
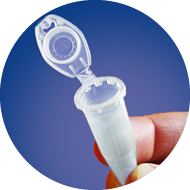
1
Only use 1.5 ml
safe-lock tubes
|

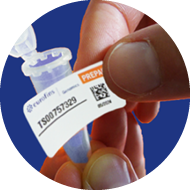
2
Remove barcode
sticker from film
|
3
Affix barcode sticker
horizontally
|
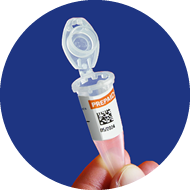
4
QR-code visible at front so
it can be read by scanner
|