Introduction
You have prepared your DNA construct, cloned it into a vector and transformed E. coli with it. Now it is time to pick and grow the colonies and conduct minipreps in order to extract plasmids to send for sequencing. Confirmation of a construct’s presence and identification of the construct in the vectors is absolutely essential before you continue to work with the plasmids. However, this process is quite laborious and time-consuming, especially when there are a large number of colonies to check.
You can save time by using the Direct Colony Sequencing solution. This Sanger sequencing service in 96-well plates conveniently sequences the plasmids/vectors directly from bacterial colonies without the need for plasmid preparation.
You just need to pick the colonies and directly transfer them to the 96-well plate that comes with the service. The wells of this plate contain buffer to preserve the colonies.
This fast and cost-effective solution can be used to quickly screen your DNA cloning experiments and select the candidate(s) for final confirmation via DNA preparation and sequencing.
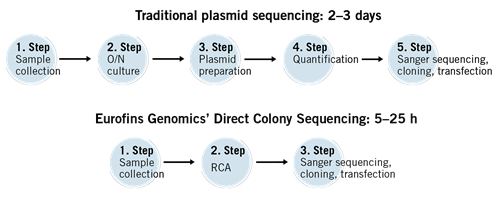
What is the basis of Direct Colony Sequencing and why does this sequencing solution work without plasmid preparation?
Rolling circle amplification process
The basis for the Direct Colony Sequencing solution is the rolling circle amplification (RCA) technique that generates multiple copies of circular DNA.
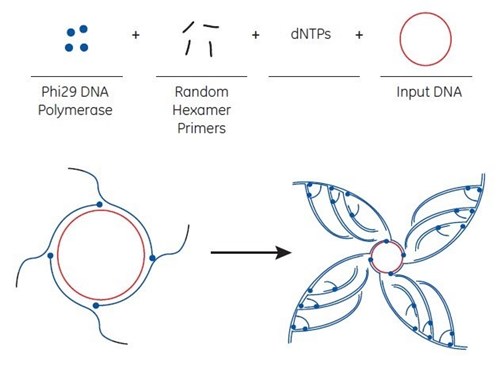
The rolling circle amplification (RCA) method utilises bacteriophage Phi29 DNA polymerase to exponentially amplify single- or double-stranded circular DNA templates. The isothermal amplification method produces microgram quantities of DNA from picogram amounts of starting material in a few hours.
An in vitro amplification of very small amounts of template DNA eliminates the need for overnight cell culture and conventional plasmid or M13 bacteriophage DNA purification. The proofreading activity of the Phi9 DNA polymerase ensures high fidelity DNA replication.
Starting material
The starting material for amplification can be of various sources, such as from a small number of bacterial cells containing a plasmid, isolated plasmids and intact M13 phages to any circular DNA (including ligation reactions).
Bacterial colonies can be picked from agar plates and added directly to the 96-well plate. Alternatively, microlitre quantities of a saturated bacterial culture or a glycerol stock can also serve as starting material.
Process
RCA is performed at 30°C without thermal cycling. Depending on the source of starting material, amplification is completed in 6 to 18 hours. The amplification products are double-stranded concatemers of the circular template with high molecular weight. Concatemers are large DNA molecules that contain many copies of a specific DNA sequence.
For RCA, random hexamer primers anneal to the circular single-strand template DNA at multiple sites. The Phi29 DNA polymerase extends each of these primers. When Phi29 reaches a downstream extended primer, strand displacement synthesis occurs. Here, the original single-stranded DNA is replaced by the newly replicated strand. The displaced strand is then available to be primed by more hexamer primer. The process results in exponential amplification.
Note: when starting with M13 bacteriophage clones, the product is double-stranded DNA and can be sequenced with forward and reverse primers. DNA amplified by the method can be used directly without any purification.
The Direct Colony Sequencing service has been validated primarily for plasmids in E. coli colonies. In principle, the method also works with large constructs (e.g. BACs) and other organisms. Contact us for pilot testing and optimisation. As a starting point, we recommend performing cell lysis or crude DNA extraction and adding 1 to 5 µL of the lysate/extract to the buffer in the 96-well plates.
References
Dean, F. et al., Genome Research 11, 1095–1099 (2001).
Lizardi, P. et al., Nat. Genet. 19, 225–232 (1998).
Estaban, J.A. et al., J. Biol. Chem. 268, 2719–2726 (1993).
Benefit from the greatest flexibility to handle your sequencing primers.
Thanks to our DNA synthesis lab, bioinformatic know how and internal IT solutions you can choose how to provide us with your sequencing primers:
- Select from standard primers on stock
Select and manage your favourite standard primers from more than 80 different validated standard primers
- Order specific sequencing primers
Specific primers can be ordered directly with your sequencing order. These primers are stored for 4 weeks and can be re-used during this time
- Enclose your own sequencing primers
You can send us your sequencing primers in 1.5ml tubes along with your samples by using our Primer Barcodes*
Please consider optimal primer conditions, concentration and volume as described below.
Optimum primer conditions:
- Primers must not contain phosphorylation or fluorescent dyes
- The optimum primer length is between 16-25 bases
- The primer melting temperature (Tm) should be 50 - 62°C
- The GC content of the primer should be 35-60%
- Ideally one G or C should be located at the 3' primer end
- The number of 3' Gs or Cs should not exceed 2 Gs or Cs
- If possible, avoid >3 identical bases in a row in the sequence
Primer concentration and volume:
- Exactly 10 pmol/µl primer concentration is required per sequencing reaction
- Each primer must have a total volume of 15 µl (double distilled water or 5mM Tris-HCl); 5 µl of primer volume is required for every additional sequencing reaction
- Concentration of primers with wobble bases must be calculated according to the following formula: nX x ConcPrimer