Overview - What is metagenomics?
Metagenomics is the study of the entire genetic content of all microbiota members in a natural habitat by utilisation of the whole genome sequencing technique. In contrast, metataxonomics utilises only 16S rRNA analysis.
The field of metagenomics centres upon direct genetic analysis of microbial genomes isolated from various environments ranging from the human gastrointestinal tract (gut microbiome) to geothermal hot springs. Metagenomics is an alternative approach to targeted amplicon sequencing in the study of uncultured microbiota. By offering direct access to the entire genetic make-up of microbial communities, metagenomics can provide valuable molecular insights into novel enzymes and biocatalysts, as well as into genomic linkages between community function and structure. The metagenomics approach serves as a powerful tool for elucidating the relationship between host-associated microbial communities and host phenotype.
Applications - What are the advantages of metagenome sequencing?
Several distinct advantages are associated with the ability to sequence the entire genomic information of all organisms present in a complex sample:
- Unbiased results free from influence of specific genomic loci
- Independence from taxonomically informative genetic markers
- Ability to study highly diverged microbes, such as viruses
- Close estimations of microbial diversity
- Detection of abundance of microorganisms in various environments
- Analysis of unculturable microorganisms
- Information on composition as well as functional capabilities of an ecosystem
- Investigation of function genes and gene clusters
Workflow - Methods & technologies for metagenome sequencing
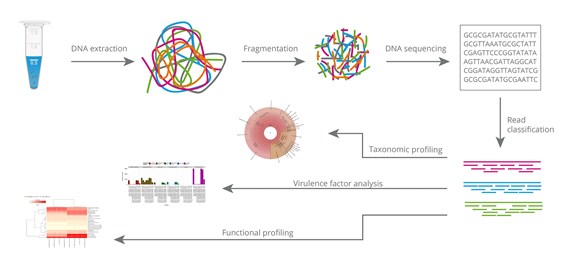
Metagenome analysis by next-generation sequencing (NGS) involves several distinct steps. Firstly, total DNA is extracted from the sample. Following a fragmentation, the DNA undergoes adapter ligation for final Illumina library preparation. The libraries are analysed using paired-end reads to maximise coverage of the amplicons. The reads are sorted and assembled into contigs. For optional de novo genome assembly, genome binning is performed with the contigs in order to reconstruct complete genomes and to assign these assembled genomes to the closest possible taxonomy. Functional analysis can be performed additionally to detect open reading frames and associated gene functions.
Genomics vs. Metagenomics vs. 16S rRNA sequencing
The main difference between genomics and metagenomics is the nature of the sample. Genomics explores the complete genetic information of a single organism only, whereas metagenomics explores a mixture of DNA from multiple organisms and entities, such as viruses, viroids and free DNA.
Amplicon sequencing, most frequently of the bacterial/archaeal 16S rRNA or fungal ITS genes, involves DNA extraction from all cells in a sample and then the targeting of a taxonomically informative genomic marker that is common to a specific group of interest. The resultant amplicons are sequenced and bio-informatically assessed to determine which microorganisms are present in the sample and in what relative abundance.
In metagenomic next-generation sequencing, DNA is also extracted from all cells in a community. However, instead of targeting a specific genomic locus for amplification, all DNA is sheared into fragments and independently sequenced by a shotgun sequencing approach. The resulting reads are aligned to various genomic locations for all genomes present in the sample, including non-microbes.
Scientific expertise: Metagenome sequencing
Eurofins Genomics has many years of experience with metagenomics analysis. The sequencing infrastructure to generate the large volume of data needed to obtain meaningful results with fast turnaround times is well in place. Eurofins Genomics has an optimised BioIT pipeline for the complex bioinformatics analysis involved in the process. Our protocols deal particularly well with metagenomes that contain unwanted host DNA. Such samples are handled with specialised techniques for selectively enriching microbial DNA prior to sequencing and with proprietary bioinformatics methods to filter host DNA and contaminant sequences during data processing.
Related products to metagenomic sequencing
Eurofins Genomics offers an all-inclusive solution for metagenomics sequencing – the INVIEW Metagenome. Molecular microbiologists can take advantage of quick delivery times and high quality data. We offer a complete package from DNA isolation to library preparation to Illumina sequencing to BioIT analysis in order to generate a comprehensive final data report for our customers.
Are you profiling bacteria with a well-defined genetic marker like the 16S rRNA gene instead? Then try our INVIEW Microbiome for in-depth characterisation of complex microbial communities.
Are you exploring a novel organism without any reference genome sequence?
If you have a limited number of samples and are only interested in a few specific genes, then see how Sanger sequencing Services can propel your metagenomics project forward.