Overview - What is epigenetics?
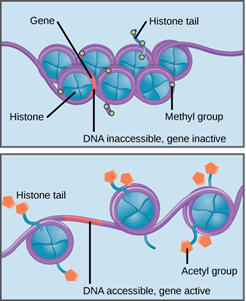
The study of epigenetics investigates modifications on DNA or DNA-associated proteins, specifically how covalent modifications affect gene expression and activity. Epigenetic mechanisms include methylation and hydroxymethylation of nucleotide bases, histone modification and regulation by non-coding RNA. Epigenetic modifications can be stable and heritable or they can be dynamic and influenced by outside factors like age and environment.

Why is epigenetics important?
Epigenetic factors regulate key cellular processes like normal development and adequate response to environmental cues. Misregulated epigenetic mechanisms can cause abnormal gene expression by improper activation or silencing of genes. These disrupted epigenetic systems often contribute to disease pathogenesis of serious disorders like cancer or neurodegenerative, developmental and autoimmune diseases.
Workflow - How can epigenetic modifications be detected?
DNA methylations and histone modifications can be investigated by using standard molecular biology methods or approaches based on next-generation sequencing.
Bisulfite treatment is a common technique that serves as the starting point for profiling human DNA methylations. In short, sodium bisulfite converts only non-methylated cytosines into uracils, leaving methylated cytosines unmodified. Downstream analysis of the bisulfite-treated DNA through PCR, next-generation sequencing (NGS) and bioinformatic analysis distinguishes between methylated and non-methylated cytosines and allows for quantitative analysis.
Several restriction digestion enzymes have target sites that are influenced by the methylation status of the genome. These restriction digestion enzymes cut DNA segments differently depending on the presence or absence of methylation at their recognition sites. This technique, combined with molecular biology methods like PCR, next-generation sequencing, Southern blots, primer extension, high-throughput liquid chromatography, and MALDI-TOF MS can be used to effectively investigate the methylation status of genes.
Methylated DNA Immunoprecipitation uses antibodies that specifically recognise methylated DNA fragments to target and isolate only methylated regions of the genome. The immunoprecipitated methylated DNA can be further analysed using micorarrays or next-generation sequencing.
Histones are the main protein component of chromatin. Histones play an important role in the regulation of gene expression by undergoing modifications like methylation or acetylation, often on their lysine or arginine residues - histone modifications. The most commonly used technique for analysing protein interaction with DNA is chromatin immunoprecipitation (ChIP). ChIP can also help to identify regions of the genome that are associated with specific histone modifications. ChIP involves crosslinking DNA-associated proteins to DNA, immunoprecipitating the DNA via protein-specific antibodies, purifying the enriched fragments and subjecting them to further analysis. Single protein-binding sites can be investigated using PCR with primers for specific genes of interest. On a genome scale, the DNA can be analysed via microarrays (ChIP-chip) or via next-generation sequencing (ChIP-seq).
Did you know that we at Eurofins Genomics have great expertise in Next generation sequencing?